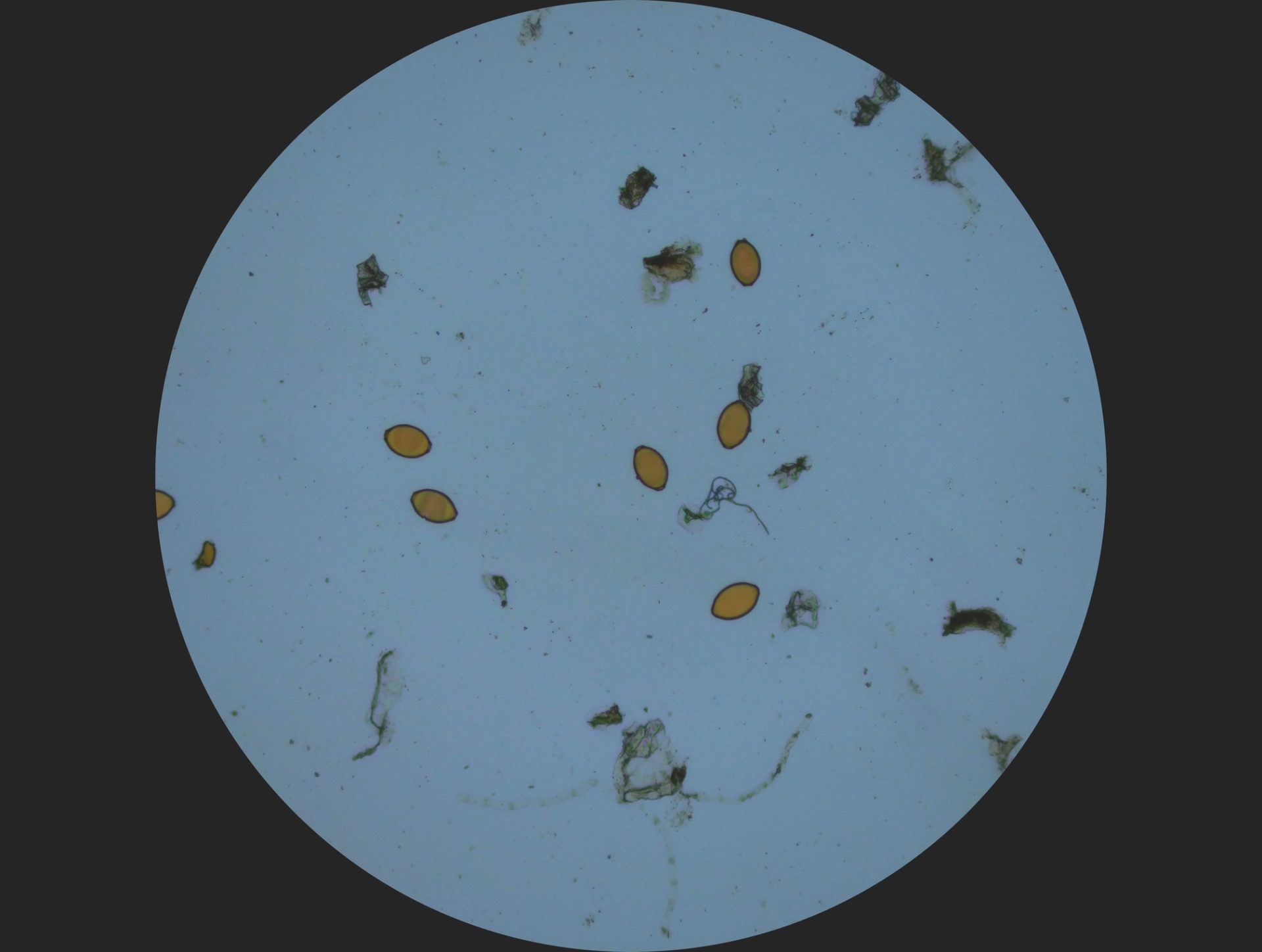
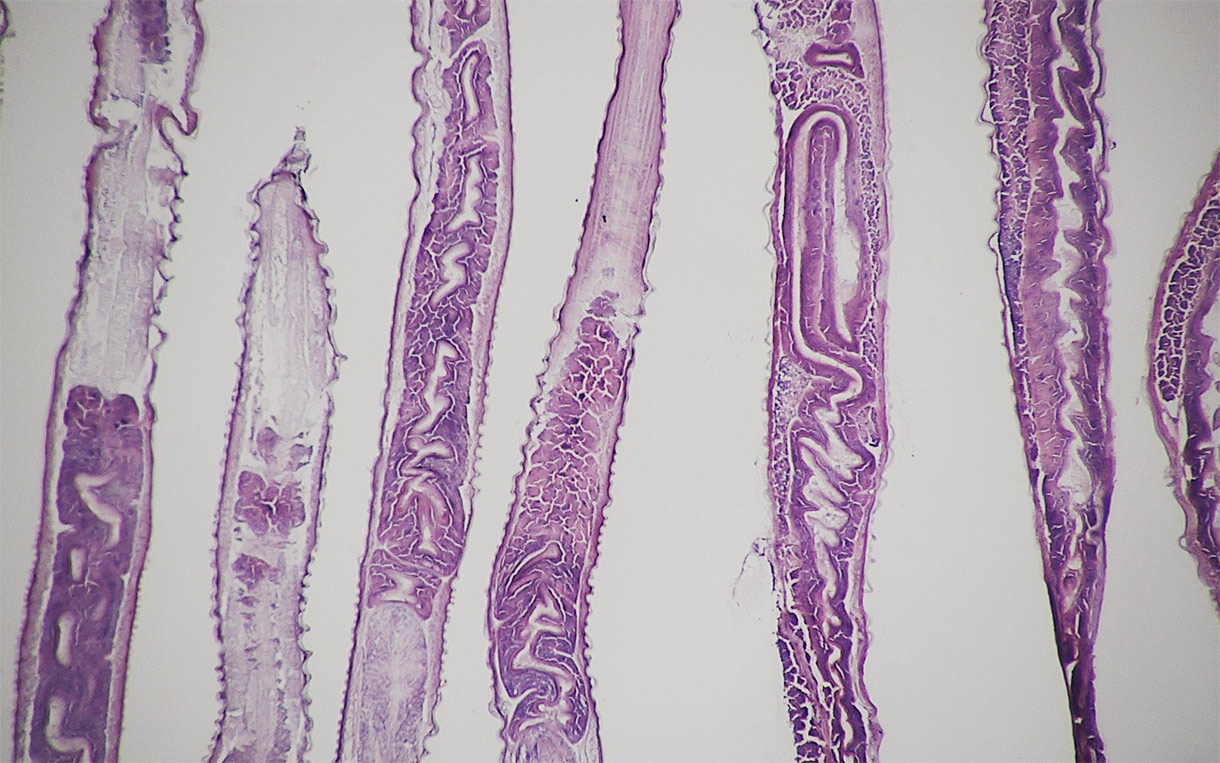
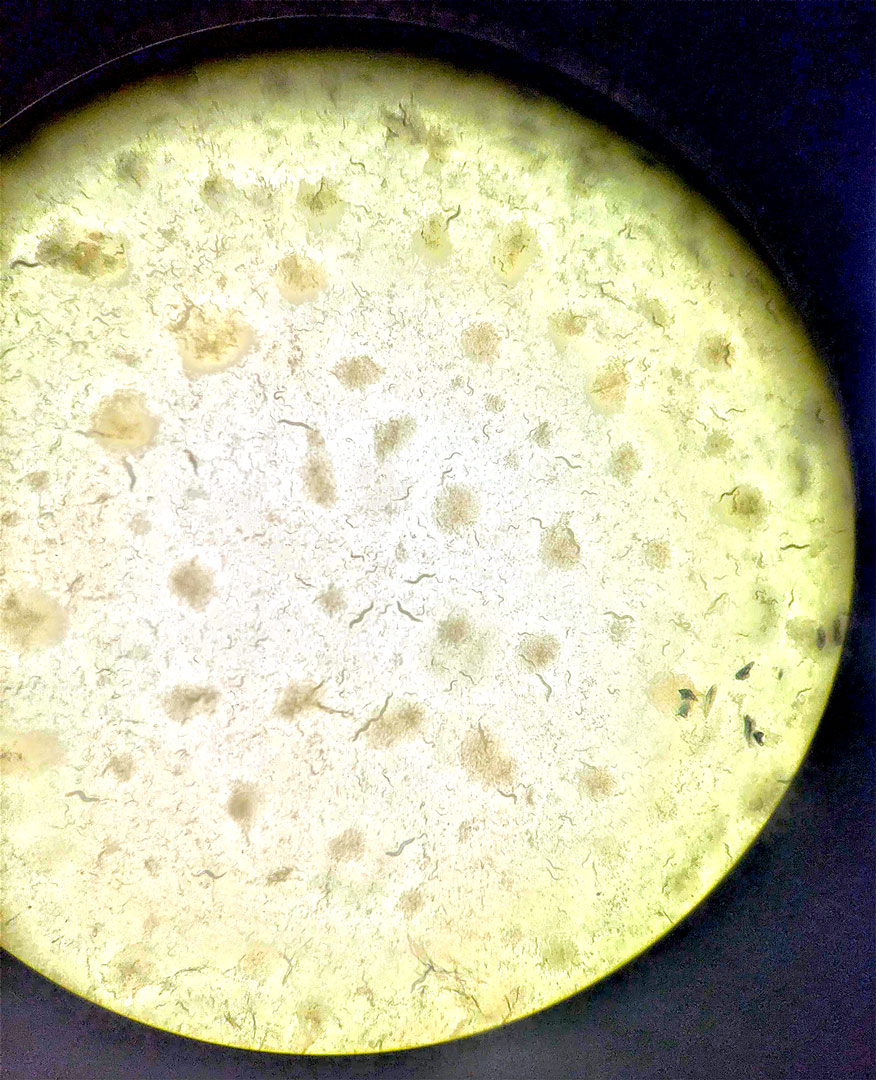
Haemonchus contortus, a parasitic worm that infects sheep, with its eggs. Image credit: Oregon State University / Wikimedia Commons.
We explore how genomics is helping researchers at the Wellcome Sanger Institute to unpick the tangled world of parasitic worms.
Look down at your skin. Imagine a tiny line, pale at first, creeping across the surface. A worm, no bigger than a thread, slipping slightly under your flesh. You did not see it happen or feel it enter – but it is in there, moving. It burrows deep down, leaving a trail that your body only notices as an itchy or crawling sensation that should not be there. Are you now itching? Apologies.
This is the hidden life of helminths – parasitic worms that live alongside us, unseen yet persistent. They can silently weaken us, causing malnutrition, disease and in severe cases, death. Beyond humans, they devastate livestock, gnawing at health and productivity, leaving farmers struggling and economies strained. These invaders, often nearly invisible in their early life stages, wield a power far larger than their size suggests.
In this blog, we dive into the world of helminths – what they are, why we study them here at the Wellcome Sanger Institute and how unravelling their biology could lead to more targeted therapies to fight these persistent parasites.
What is a helminth?
The definition of a helminth depends on which expert you ask. The term helminth itself comes from the Ancient Greek word ἕλμινς (hélmins), which means worm.
In the early days of parasitology, scientists would open a human or animal and simply notice worms inside. The reaction was straightforward: “There are worms here. Let us call them helminths.” There was no concern for evolutionary relationships – these worms were grouped together because they shared one thing: they parasitised a host.
From a modern phylogenetic perspective, helminths are not a natural taxonomic group but a practical umbrella term for parasitic worms drawn from several distantly related animal lineages. These include parasitic flatworms (Platyhelminthes), nematodes (Nematoda) and acanthocephalans (Acanthocephala), among others. Importantly, parasitism has evolved multiple times within these groups, and many members of Platyhelminthes and Nematoda are free-living and play neutral or beneficial ecological roles. Humans can be infected by over 300 different species, although a much smaller number are associated with causing disease worldwide.1
Why study helminths?
Nematodes are the most abundant animal on Earth. Nathan Augustus Cobb, known as the ‘father of nematology’, said that if everything except nematodes disappeared, the Earth would still be faintly recognisable, with its mountains, hills, valleys, rivers, lakes and oceans outlined by a layer of nematodes. Adding all the other helminths on top, it becomes clear just how numerous these creatures are – showing why studying them is essential to understanding life on Earth.
Almost all vertebrates can be infected by at least one species of helminth at some point, though not continuously. In humans, helminths are part of a group of organisms that cause neglected tropical diseases (NTDs). These are a group of conditions that are associated with significant health, social and economic implications. NTDs are most prevalent in low- and middle-income countries, particularly in tropical regions. In some areas, infection rates are extremely high, disproportionately affecting children. These infections can make children feel very ill, prevent them from attending school and even impact their biological development. They can cause or worsen malnutrition, creating a continuous cycle of health and social challenges for affected communities. Estimates suggest that NTDs affect more than one billion people worldwide.2

Parasitic worms such as Haemonchus contortus cause millions of pounds' worth of damage to the global livestock farming industry. Image credit: AdobeStock.
Helminth infections are a major global constraint on livestock, reducing growth, reproduction, milk and wool production, and overall productivity. They cause diarrhoea, anaemia, increased susceptibility to other diseases, and can be fatal in severe cases. Economically, they are a significant burden – costing the UK livestock industry around £270 million annually, with drug resistance contributing £7.5 million of this.3
Treating helminth infections in both humans and animals is increasingly difficult due to rising anthelmintic resistance, which is already widespread in livestock and represents a growing risk in human populations. Many drugs are used to control helminth infections, with mass drug administration implemented across large parts of the world. In humans, this typically targets children and women of reproductive age, even if they are not infected. Many of the same classes of drugs are also used in livestock, often with less regulation. As a result, several important species of veterinary helminths have rapidly developed resistance, raising concerns that similar high rates of resistance could occur in human helminths.
Understanding basic biology and genetics
Understanding the basic biology of helminths is important because it underpins effective prevention, diagnosis and treatment of helminth infections. This in turn enables better disease control, drug and vaccine development, and reduction of their global health impact.
At the Sanger Institute, Dr Steve Doyle, Career Development Fellow, and his team in the Parasites and Microbes programme study a gut parasite of sheep called Haemonchus contortus, also known as the barber’s pole worm. It is a parasitic, blood-sucking roundworm that infects sheep, goats and occasionally other ruminants found in temperate regions worldwide. Its name is derived from its distinct red striping of the female worm as its intestine and uterus intertwine. Infection results in anaemia and possible death, causing a significant burden on the farming community due to production losses and the cost of controlling them. In the UK, the barber’s pole worm is estimated to affect around 56 per cent of sheep flocks.4
Steve and his team have done a lot of work on the parasite, including mapping the genome, and conducting genome-wide association studies to map key genes and mutations causing drug resistance worldwide.5,6
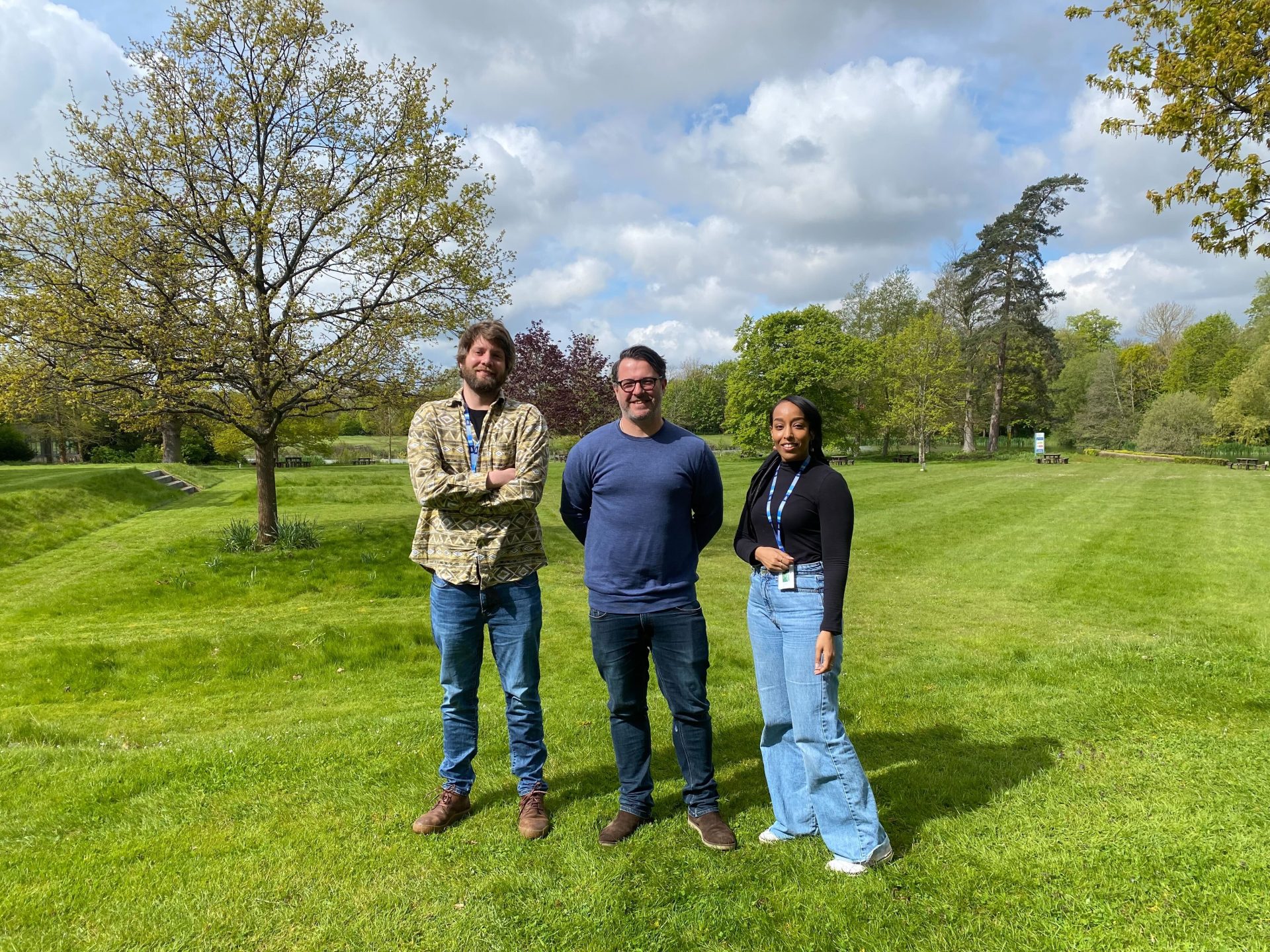
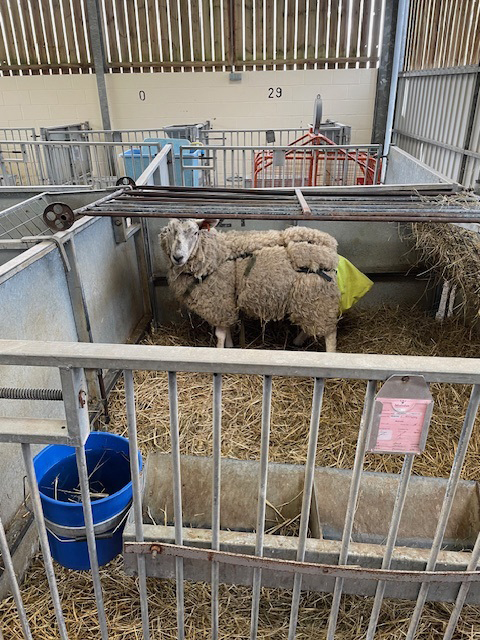
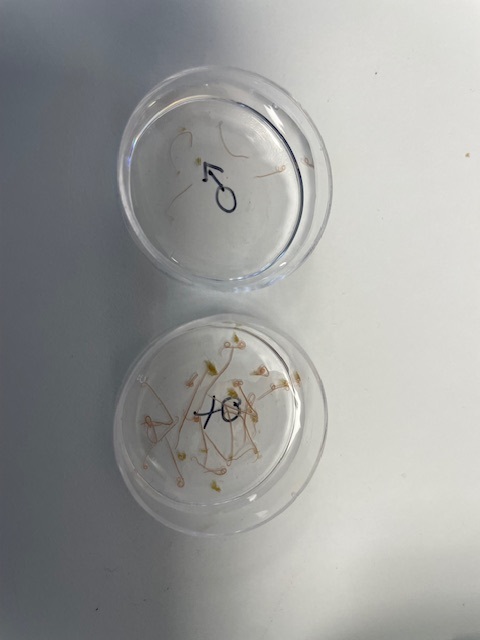
Top: The Doyle research group. Bottom left to right: Collecting worms from sheep, male and female Haemonchus contortus worms, and counting H. contortus eggs under the microscope. Images credit: Steve Doyle / Wellcome Sanger Institute.
For over a decade, the team has been working with the Moredun Research Institute in Scotland to obtain parasite samples from sheep. Because this is a natural host-parasite relationship, the barber’s pole worm is relatively easy to study compared with other species, for example, those that infect humans. Certain stages of its life cycle can be maintained in vitro, enabling researchers to conduct drug screening. Another important advantage is that samples can be frozen, allowing the team to preserve invaluable parasite isolates.
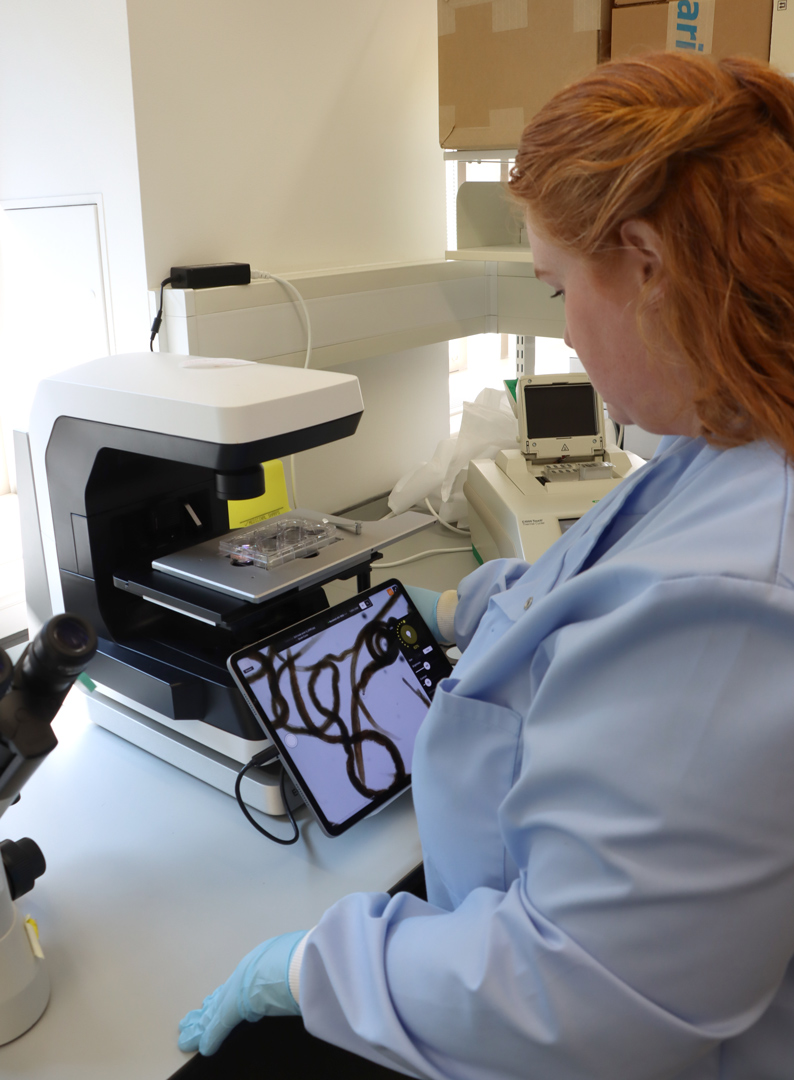
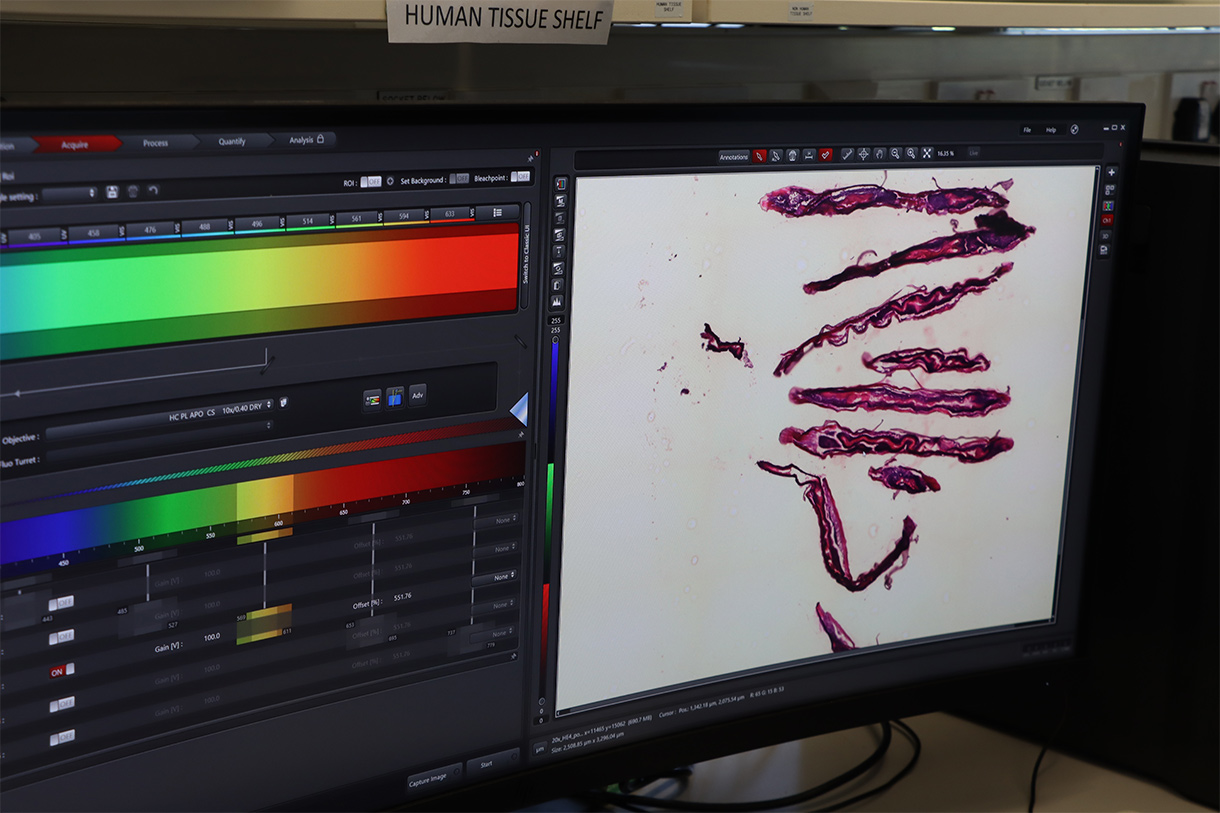
Using these samples, the team has more recently conducted extensive single-nucleus RNA sequencing – a method to analyse which genes are active inside individual cell nuclei of a tissue – to generate cell atlases of whole life stages to gain a clearer picture of the worms’ biology. Dr Sarah Buddenborg, a former Postdoctoral Fellow in Steve’s team, has also been using spatial transcriptomics to identify the different cell types present and determine which genes are active throughout the parasite's life cycle.
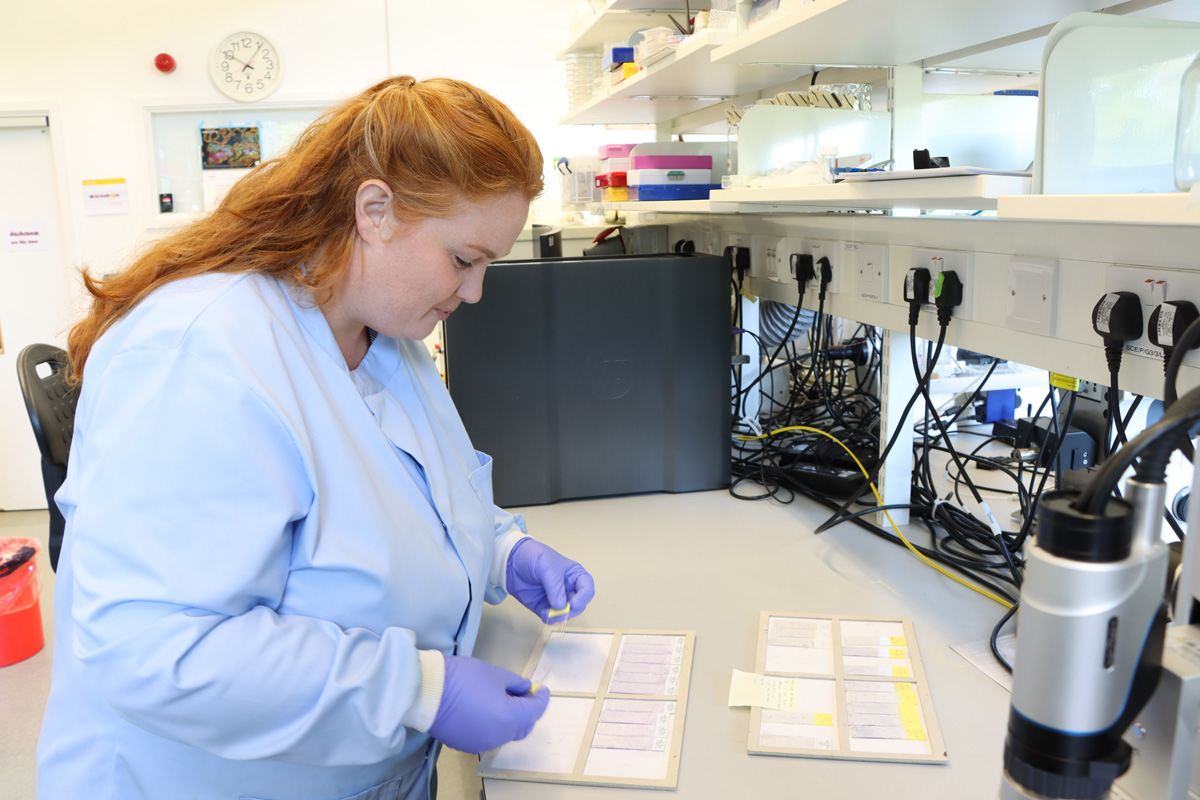
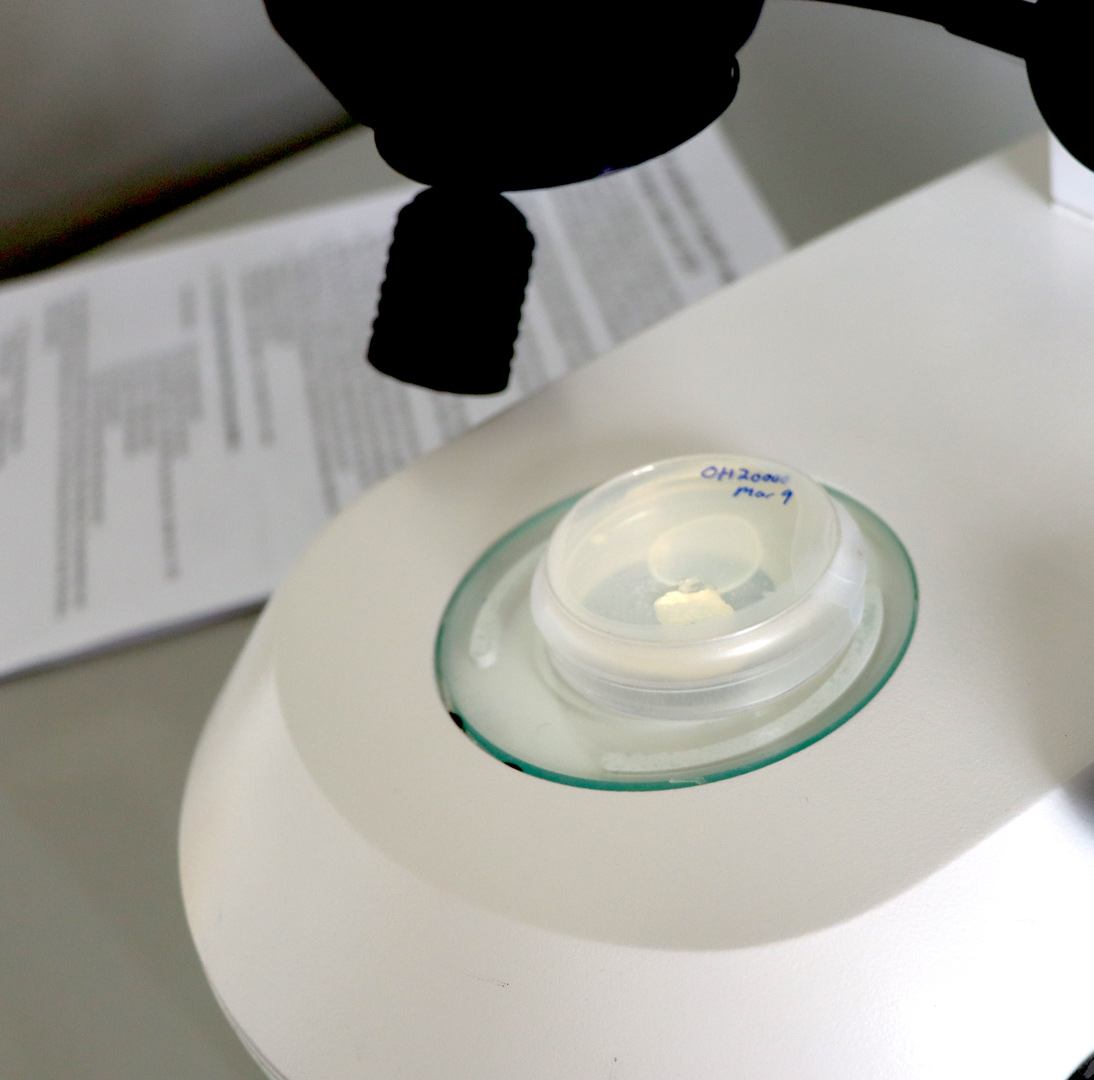
Sarah Buddenborg studying Haemonchus contortus, also known as barber’s pole worm because of its red and white stripes. The worms were too small to generate meaningful spatial genomic data, until now. Images credit: Carmen Denman Hume / Wellcome Sanger Institute.
“Helminths exploit an incredible array of strategies to efficiently and effectively parasitise their hosts. They have been so successful, in part, because of their genetic diversity, which also enables them to rapidly evolve resistance to drugs used to control them. The cell atlases generated in our group are providing amazing insights into their biology and will almost certainly reveal hidden vulnerabilities that could be exploited for new control options. Further, our genomics work on drug resistance variants has begun to redefine our understanding of the genetic mechanisms by which parasites become resistant, and offers real promise for the development of genetically-based diagnostic tools to better manage parasites and improve animal health.”
Dr Steve Doyle
Career Development Fellow, Wellcome Sanger Institute
Sequencing and studying the genomes of worms
Historically, one of the difficulties with studying parasites was that while some of them are large, the majority of them are tiny – only a few millimetres in length – and as a result, too small for standard sequencing approaches.
Thankfully, former Sanger Institute Postdoctoral Researcher and now collaborator at the Natural History Museum, Dr Chris Laumer, developed PiMmS – picogram input multimodal sequencing – to help take tiny amounts of DNA, amplify it and then sequence it using long-read platforms. Using this approach, Dr Lewis Stevens, Postdoctoral Fellow, and his colleagues in the Tree of Life programme at Sanger have been getting worm samples from up and down the country to generate genomes of parasites that were not possible to obtain before. From parasites from soldiers returning home to parasites of bumblebees, moles and hedgehogs, this work is part of a project called the 959 Nematode Genome Project, which is trying to build a massive dataset of nematode genomes.
RELATED BLOG
Ways Sanger is applying spatial transcriptomics to our science
We are harnessing spatial transcriptomics to uncover how cells behave within their natural environments, whether in the human skin or in a parasitic worm. Collaborations across the Institute between our scientific programmes and our operational support teams are enabling us to push boundaries and provide insights that will transform our understanding of basic biology.
By sequencing genomes from individual worms, Lewis can now compare parasites at a much finer scale than was previously possible. This has revealed regions of the genome that differ markedly between individual worms known as hyperdivergent haplotypes. This previously hidden layer of variation may help explain how these parasites evade the host immune system. Importantly, it also has implications for vaccine development: if proteins targeted by vaccines differ between worms, this variation could allow them to evade treatment.
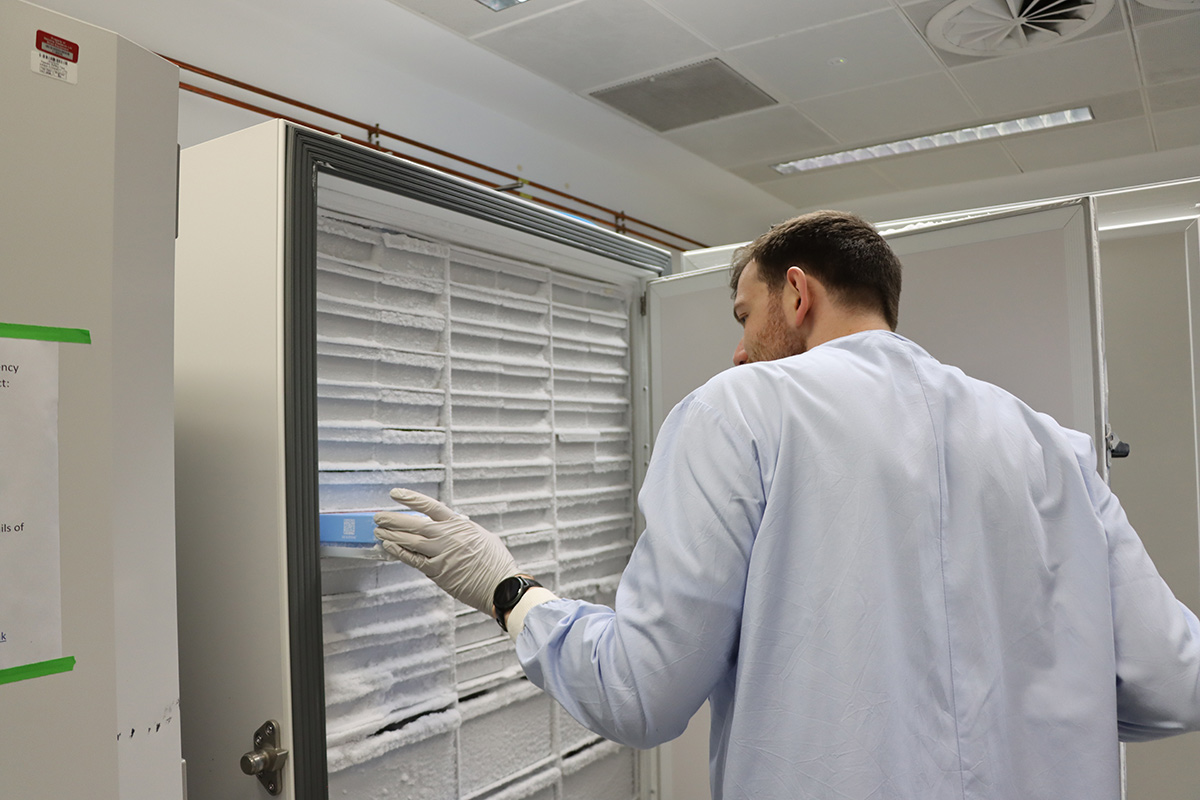
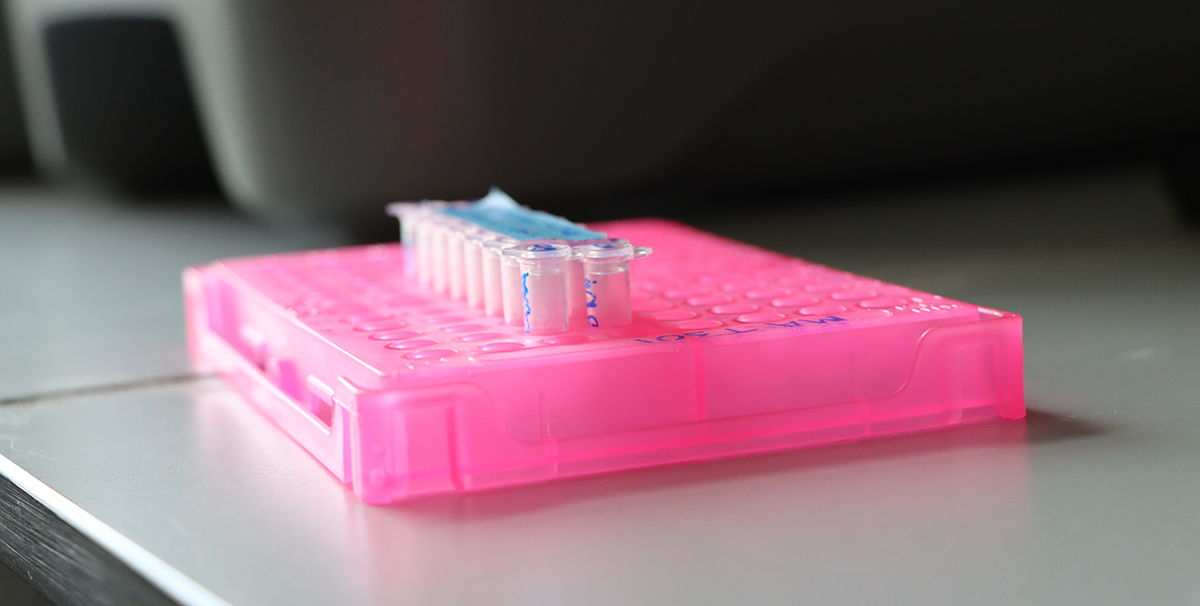
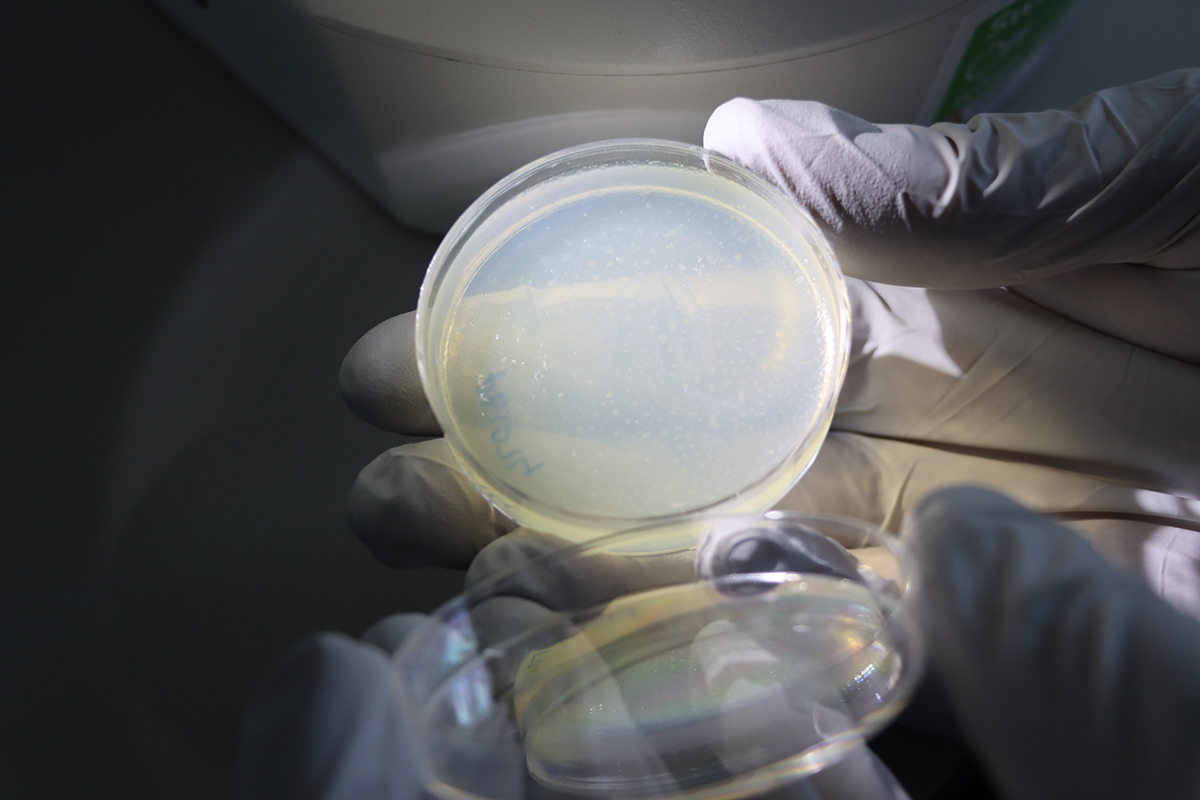
Lewis Stevens is sent samples from around the world, taken from animals infected with parasitic worms. The samples arrive at Sanger and are stored in the freezer until they are ready to have their DNA extracted for sequencing. Caenorhabditis elegans – the iconic model nematode and a free-living relative of the parasites Lewis studies – can be seen under a low-magnification microscope grazing on bacteria on an agar plate. Images credit: Carmen Denman Hume / Wellcome Sanger Institute.
Another piece of work that Lewis has been exploring is whether we can obtain complete genomes from stool samples. Most genome sequencing efforts to date have focused on adult parasites, which typically must be collected from dead hosts, making sequencing difficult or impossible in many systems. Therefore, with funding from the Sanger Postdoc Committee, Lewis has been piloting a study to explore an easier approach for sampling. He obtained some stool samples from the remote islands of St Kilda, Scotland, where feral sheep roam the land – and now act as a long term study system for researchers interested in evolution, ecology and climate change.
Presented with some murky water, Lewis pulled out the individual worms, brought them back to Sanger and sequenced them in our labs. From this, Lewis was able to generate a complete genome from the stool sample – demonstrating that stool is an accessible alternative to dissecting dead hosts.
“Genomes are key to understanding how these parasites work, but until now they’ve been difficult to sequence, meaning we only have high-quality genomic data for a handful of species. The methods we’re developing at Sanger make it much easier to generate parasite genomes, even directly from faeces. This will dramatically expand the number of species we can study, enabling everything from cross-species comparisons to analyses of variation between individual parasites.”
Dr Lewis Stevens,
Postdoctoral Fellow, Wellcome Sanger Institute
Tracking resistance to evaluate new treatments
Another part of Steve Doyle’s research, alongside Staff Scientist Dr Javier Gandasegui in his group, is identifying effective treatments against soil-transmitted helminths. These are a group of common intestinal parasite worms that spread through soil contaminated with human faeces. They affect around a quarter of the world’s population.7
Steve and Javier are part of a consortium called STOP – Stopping Transmission of intestinal Parasites – which unites institutes across Europe and sub-Saharan Africa to find a more effective treatment against soil-transmitted helminths. In 2021, the consortium began a Phase 3 clinical trial which explored a new drug formulation, consisting of the standard drug, albendazole, with a different drug called ivermectin. The safety and efficacy of this was tested in 1,250 school children from Ethiopia, Kenya and Mozambique.
In 2023, the Sanger Institute joined the second consortium called STOP2030, where Steve is a co-investigator on the grant that funds the work. This consortium will build off the previous, running a larger Phase 4 trial to evaluate the safety of this coformulation across 20,000 school children from Ghana and Kenya. The hope is that these findings will be used to inform the World Health Organization’s recommendation for this new drug in mass drug administration campaigns.
Steve’s team at the Institute will be involved in generating genome wide data from the parasites to see if they can identify mechanisms of resistance. The team is also studying how the microbiome can influence drug efficacy and safety, and the impact of anthelmintic treatment – drugs used to treat helminths – on the gut microbiome.
“I work on neglected tropical diseases, which are very important to study, because we have real capacity of generating evidence that can positively impact the most vulnerable communities in the world. I believe with the right level of investment these infections can be eliminated easily. The first disease to be eradicated on Earth was smallpox and I think that the next one will be the helminth, Guinea worm, which causes Dracunculiasis – a debilitating parasitic infection. These parasites are ancient, but our tools are new – and for the first time, eradication is not a distant ideal but a realistic possibility.”
Dr Javier Gandasegui
Staff Scientist, Wellcome Sanger Institute
References
- Owusu-Agyei AK, Nabarro L. Helminth infections and differentials of eosinophilia. Medicine. 2021; 49: 766–769. doi: 10.1016/j.mpmed.2021.09.008
- World Health Organization. Neglected tropical diseases [Last accessed: February 2026]
- Charlier J, Rinaldi L, Musella V, et al. Initial assessment of the economic burden of major parasitic helminth infections to the ruminant livestock industry in Europe. Preventive veterinary medicine. 2020; 182: 105103. doi: 10.1016/j.prevetmed.2020.105103
- Rare Breeds Survival Trust. Silent Killers: Barber pole worm (Haemonchus contortus) [Last accessed: February 2026]
- Doyle SR, Tracey A, Laing R, et al. Genomic and transcriptomic variation defines the chromosome-scale assembly of Haemonchus contortus, a model gastrointestinal worm. Communications biology. 2020; 3: 656. doi: 10.1038/s42003-020-01377-3
- Doyle SR, Laing R, Bartley D, et al. Genomic landscape of drug response reveals mediators of anthelmintic resistance. Cell Reports. 2022; 41: 111522. doi: 10.1016/j.celrep.2022.111522
- World Health Organization. Soil-transmitted helminth infections. January 2023 [Last accessed: February 2026]